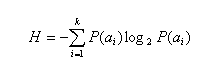
where: {a1, ..., ak} is the alphabet of the size k, and P(ai) is a fractional composition of ai
Program for mapping low complexity regions in protein sequences. Search for the low complexity regions is performed with using Shannon's information measure.
Search for the low complexity regions is performed with using Shannon's information measure. Shannon's information is defiened as follows:
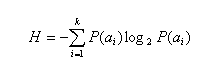
where:
{a1, ..., ak} is the alphabet of the size k, and P(ai) is a
fractional composition of ai
The search is carried out as follows. For each position i of the sequence S calculation of the Shannon's information H(i, l) is performed in the window of size l within the range [lbegin, lend]. If H(i, l) turns out below prespecified threshold Hthr(l) then fragment [i, i+l] is declared low complex. Intersection of all such fragments at the end of calculation gives a map of low complexity regions of the sequence S.
>EXAMPLE SEQ Masked regions: p1: 81 p2: 120 l: 40 chain(+) [Low Complexity Region] p1: 81 p2: 120 l: 40 chain(+) [Low Complexity Region] p1: 81 p2: 120 l: 40 chain(+) [Low Complexity Region] p1: 81 p2: 120 l: 40 chain(+) [Low Complexity Region] ....
>EXAMPLE SEQ ASFDPHEKQLIGDLWHKVDVAHCGGEALSRMLIVYPWKRRYFENFGDISNAQAIMHNEKVQAHGKKVLASFGEAVCHLDG XXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXIRAHFANLSKLHCEKLHVDPENFKLLGDIIIIVLAAHYPK DFGLECHAAYQKLVRQVAAALAAEYHIGDLXXXXXXXXXXXXXXXXXX ....
>EXAMPLE FILE asfdphekqligdlwhkvdvahcggealsrmlivypwkrryfenfgdisnaqaimhnekvqahgkkvlasfgeavchldg EEEEEKKKKKEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEirahfanlsklhceklhvdpenfkllgdiiiivlaahypk dfglechaayqklvrqvaaalaaeyhigdlEEEEEEEEEEEEEEEEEE ....